Feature
Upping our game
Multidrug-resistant infections already kill five million people each year. Will new discoveries put science back on top?
By Joshua Krisch Illustration by Ibrahim RayintakathOne day, Elizabeth Campbell got a call from a group of frustrated researchers at the University of Zurich. The Swiss scientists were trying to coax the antibiotic fidaxomicin—commonly used to treat infections from C. difficile (a germ that typically attacks the large intestine, causing colitis and other problems)—to beat back drug-resistant tuberculosis. Fidaxomicin had been shown to do a great job of killing Mycobacterium tuberculosis cells in a petri dish; the researchers couldn’t figure out how to move the drug from the gastrointestinal tract into the bloodstream and on to the lungs, where TB lives.
Colleagues often turn to Campbell for help with such problems. Maybe they’ve seen promising experimental results for a potential new drug but can’t explain the mechanism of action. Or maybe they’ve unexpectedly hit an impasse in a once promising line of experiments. In the eyes of her peers, Campbell, a structural biologist in Rockefeller’s Laboratory of Molecular Biophysics, is something of an antibiotics whisperer.
The COVID pandemic accelerated what had already been a crisis, creating a petri dish for breeding drug resistant bacteria.
“We analyze, step-by-step, how potential drugs behave inside the cell—how an antibiotic attacks a bacterium, for instance,” says Campbell, who is also a research associate professor. “In the case of fidaxomicin, we were able to suggest a few specific adjustments that could potentially nudge the drug into the bloodstream.”
It’s hard to overstate how precious this type of information could be. TB is no longer the disease it was in the latter half of the 20th century, following the 1943 discovery of streptomycin, a potent antibiotic. Virtually overnight, a disease that had terrified humanity for millennia—responsible by the 1800s for, and more lethal than the Black Death, leprosy, or AIDS—was finally being tamed.
Today, more than 80 years later, multidrug-resistant TB strains kill hundreds of thousands of people each year. And what keeps researchers like Campbell up at night is the knowledge that this emerging resistance could someday render the disease impossible to treat once again—making diagnosis akin to a death sentence.

What’s more, TB is just one potential tidal wave in the hurricane of antibiotic resistance headed our way.
“We’re in a state of constant warfare with all infectious microbes,” Campbell says. “Every time we innovate, they immediately respond by mutating.”
At Rockefeller and beyond, scientists like Campbell are tenaciously working to keep medicine one step ahead. They’re leveraging the latest advances in AI and CRISPR to reveal new genetic targets, learning new techniques from the age-old battle among microbes, and opening up entirely new approaches by unpacking the basic biology of their microscopic foes.
The stakes could not be higher. Whereas the golden age of antibiotic discovery—a period that stretched from the 1930s through the 1950s—miraculously reduced the lethality of infections ranging from TB to rheumatic fever, so-called superbugs that can shrug off any drug we throw at them now kill someone in the United States every 15 minutes. Globally, five million people die from drug-resistant bacterial infections every year, a number expected to double by 2050. The World Health Organization has declared that if left unchecked, antibiotic resistance will pose one of the greatest threats to us as a species.
So how did a signature advance of the 20th century become an existential crisis of the 21st?
The economic toll of a post-antibiotic age
The human costs of antimicrobial resistance (AMR) are easy to see—deadly diseases run rampant, treatments grow scarce, and death rates soar. But the economic toll of AMR may also trigger an unprecedented global recession—or even spell the collapse of global trade.
The economic toll of a post-antibiotic age
HEALTHCARE

![]()
$1.5 BILLION
Cost of developing an antibiotic in the United States. The average revenue per year for a new antibiotic is estimated at $46 million, one reason why so few are in the pipeline.
Illustrations by Dan Matutina
By 1924, four years before Alexander Fleming discovered penicillin, doctors had already documented resistance to the first antibiotic (arsphenamine, a cure for syphilis). Even then, scientists saw the basic outline of the problem: The bacteria most susceptible to an antibiotic are killed, leaving only those that are resistant to proliferate. What’s more, because bacteria multiply quickly, they quickly acquire random mutations—and the quicker they reproduce, the higher are their odds for picking up resistance-conferring mutations. Those variants can then quickly spread as bacteria readily exchange DNA with one another. The COVID pandemic accelerated what had already been a crisis of growing resistance, as millions of patients were admitted to hospitals where they often received treatment for secondary infections while already immunocompromised—creating a global petri dish for breeding drug-resistant bacteria.
What’s at stake isn’t just our ability to treat infections but also a staggering number of other advances in modern medicine that depend on easy access to powerful antibiotics. When these drugs are no longer effective, routine surgeries—appendectomies, hip replacements, and nearly all elective procedures—will often be ruled too risky. Cancer treatment will be set back decades (chemotherapy ravages the immune system), and childbirth will end more often in tragedy than in celebration.
Countering resistance has always meant devising new treatments. But when it comes to antibiotics, precious few are being discovered. Between 1909 and 1990, 110 antibiotics came to market. From 2017 to 2021, just a single drug capable of treating the most critical class of resistant bacteria had been approved, and today only two more such drugs are in clinical trials. Currently, 27 antibiotics for the most threatening infections are undergoing trials. The pipeline has essentially run dry.
The reasons are complicated. For starters, as is the case with many modern medicines, it can cost hundreds of millions of dollars to develop and test new antibiotics—there are new targets to identify, refinements to add, and clinical trials to run. But unlike other desperately needed drugs, new antibiotics are designed not to be used until absolutely necessary, and even then as sparingly as possible, significantly slowing the return on investment. This has led many pharmaceutical companies to deprioritize such research.
Because approved drugs don’t have to revisit that lengthy process, advances in the world of antibiotics have recently centered around improving existing compounds—a strategy in which Campbell’s ability to pinpoint a new receptor or other molecular subtlety can mean the difference between life and death. But this is just one piece of the puzzle.
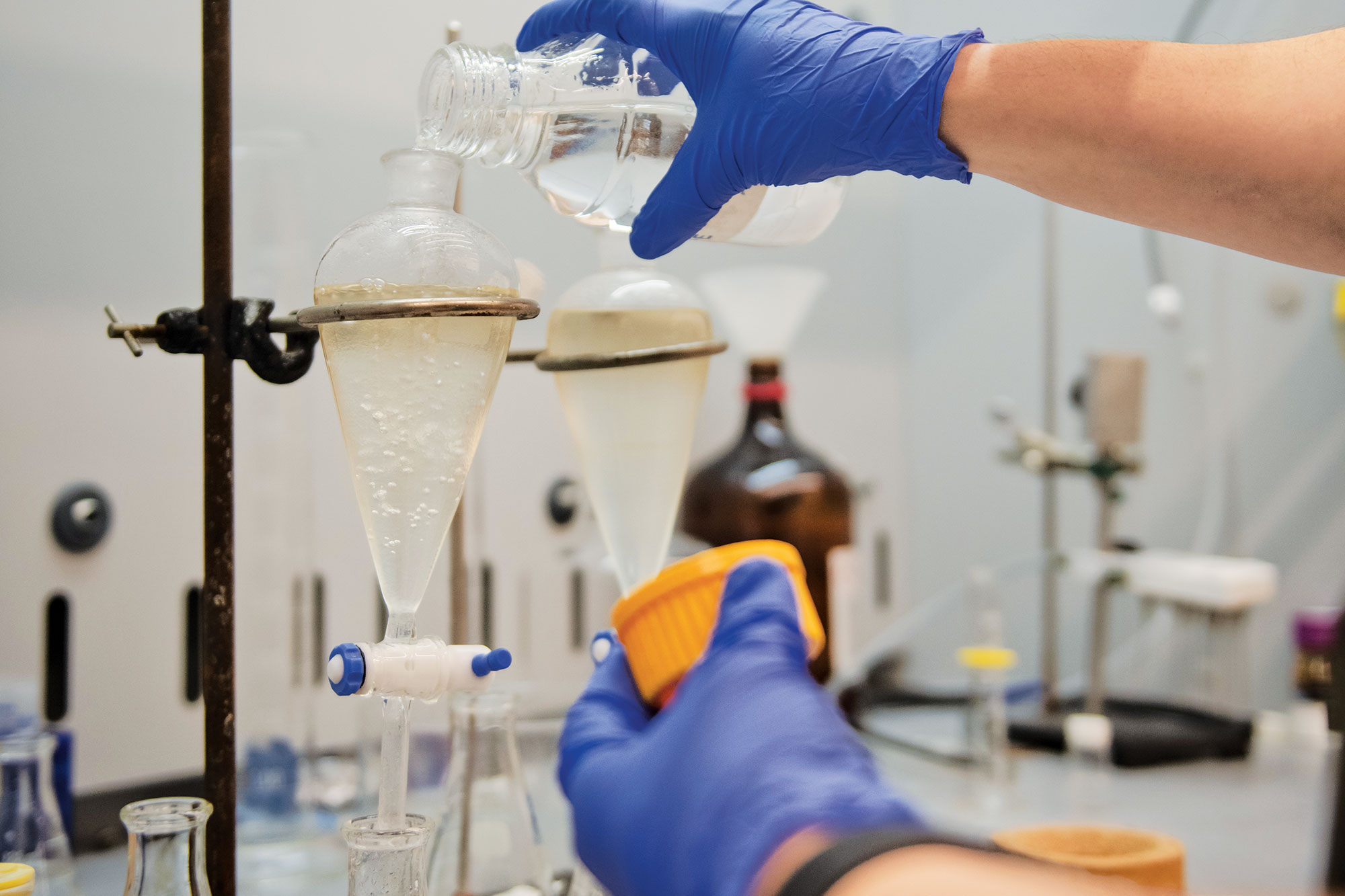

When scientists talk about new antibiotics, they mean something truly novel: a compound that looks like nothing present-day bacteria have seen before. That is exactly what Sean F. Brady, Rockefeller’s Evnin Professor and head of the Laboratory of Genetically Encoded Small Molecules, hopes to mine from dirt.
Brady began this work in graduate school, where he coaxed fungi into producing the small molecules that serve as the basis for most pharmaceuticals. Now, he is trying to convince soil bacteria to manufacture the very antibiotics that could be their undoing.
With enough alternatives on hand, countering resistance could come down to a numbers game.
It’s not an entirely original idea. Streptomycin was derived from soil bacteria, and today most antibiotics on the market are similarly taken directly from microbes or synthesized in the lab to resemble natural bacterial products. Bacteria have been in a mutual arms race for eons, evolving ways to kill off microscopic neighbors that encroach on their territory. There is nothing better at killing bacteria than other bacteria, and the heyday of antibiotic drug discovery more or less revolved around scientists collecting disparate samples from near and far, growing Streptomyces or Bacilli in the lab, and bottling their secrets.
Today scientists know that bacteria store the genes that produce their chemical weapons in biosynthetic gene clusters—families of about 40 genes that encode natural antibiotics—and that most of these clusters cannot be expressed in the lab. “There’s all this hidden information in gene clusters that we never see,” Brady says, “books we can’t read that would tell us how to make new antibiotics from scratch.”
His lab focuses on unlocking these mysterious gene clusters in hopes of spawning an entirely new cache of drugs that bacteria have not yet learned to resist. One of Brady’s methods involves using detailed bioinformatic analyses to tease apart the genetic instructions within a DNA sequence in order to predict the structure of any antibiotic-like compounds that a bacterium with those sequences might produce. Synthetic chemists can then take that data and synthesize a new drug candidate.
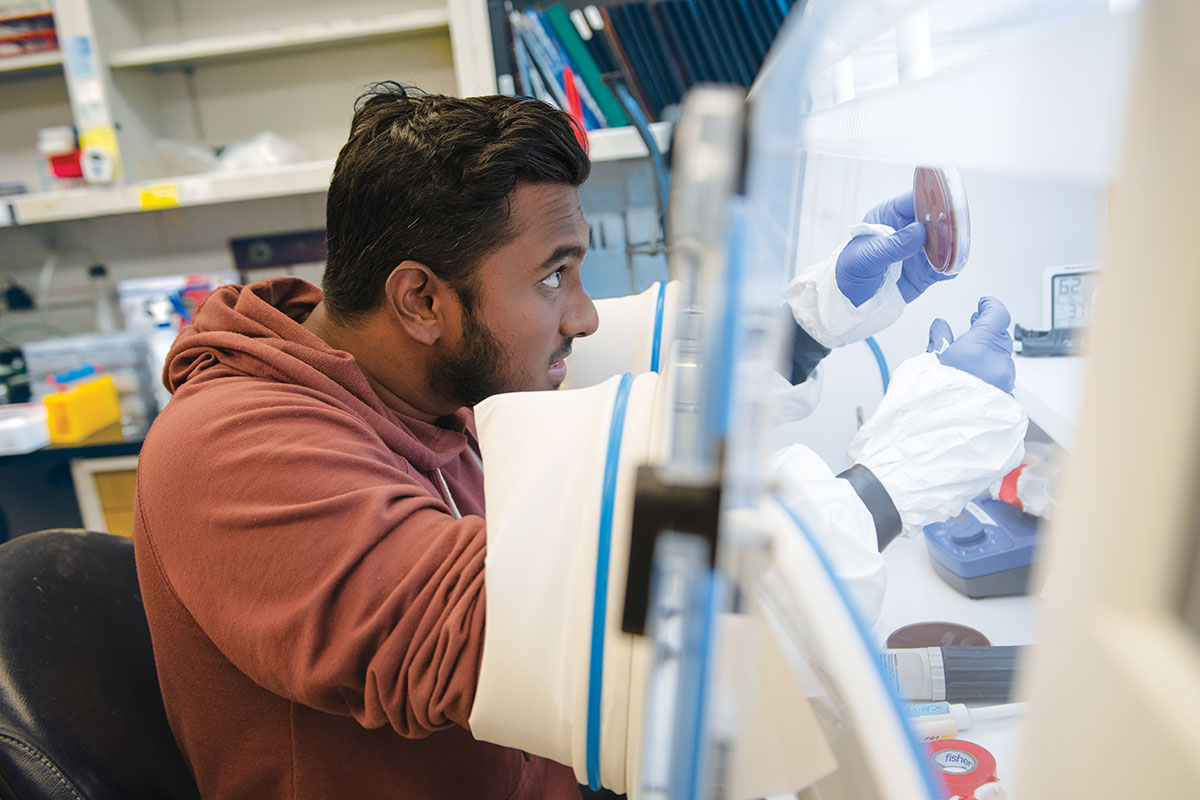

“We felt the field had gotten to the point where informatics tools were powerful enough to predict a molecule from a gene cluster,” says Brady, who refers to the technique as bioinformatic prospecting. “We were right, and that has led to a growing number of antibiotics being produced in a biology-independent way. Computers predict it, and a synthetic chemist produces what the computers predicted.”
Several synthetic antibiotics produced by Brady in this manner are now in preclinical development, including cilagicin, which the lab discovered in 2022. Brady and his team fed the relevant bacterial DNA sequences they had identified into an algorithm and synthetically produced the compound, which is effective against several drug-resistant bacteria in mice. Because cilagicin seems to disable two molecules key to keeping bacterial cell walls together, it may prove more potent than existing antibiotics. Resistant bacteria can adapt to life without one of those two molecules but have not yet learned to cobble together a cell wall when both molecules are offline.
One day, though, they probably will. But with enough alternatives on hand, countering resistance could come down to a numbers game. For instance, if we were to place penicillin on the back burner for 30 years while a drug like cilagicin took center stage, penicillin resistance might naturally fade away. And with a sufficient number of options to cycle through, humanity may be able to keep bacteria on their toes indefinitely.
The economic toll of a post-antibiotic age
PRODUCTIVITY
![]()
2 YEARS
Many laborers will die younger. As a result, the world’s working-age population is expected to decrease by two years in the next decade.
The world’s largest collection of strep bacteria sits in Vincent A. Fischetti’s lab. Fischetti runs Rockefeller’s oldest infectious disease lab, where an ongoing battle against strep, or Streptococcus pyogenes, has been waged for nearly 100 years. To control this pathogen and other instances of antibiotic resistance, Fischetti’s Laboratory of Bacterial Pathogenesis and Immunology focuses on developing alternative treatments like lysins, enzymes produced by bacteriophages, viruses that infect bacteria. Once phages reproduce inside a cell, they use lysins to break down the bacterial cell wall and release their progeny.
The idea of leveraging whole phages rather than just their lysins for medicinal purposes has been around for a while. At about the same time that Fleming was puzzling over penicillin, other scientists were exploring the possibility of therapies that pitted phages against germs. It was an intriguing premise that lost momentum due to bad timing. Once antibiotics proved so stunningly successful, Fischetti says, “whole-phage therapy was largely scrapped.”
Lysin therapy may one day prove to be one of the most durable approaches to killing bacteria.
The rise of antimicrobial resistance has changed that. There is now an entire center dedicated to phage applications at the University of California, San Diego, and the broader movement has seen some tangible success: Currently a boutique treatment with cocktails customized for individual patients, phage therapy has proven itself effective against multidrug-resistant bacteria on a limited basis.
Fischetti notes that phage therapy has been difficult to scale and that bacteria become resistant to phages very rapidly—even more rapidly than to antibiotics—because just as phages evolved to target bacteria, bacteria have evolved to evade phages.
The lysin therapy approach he uses, by contrast, can be scaled like current antibiotics. More importantly, no resistance has been seen for lysins during the 20 years of their development.
Initially, Fischetti used lysins exclusively as a laboratory tool. He purified the enzymes for his Ph.D. thesis and used them to extract material from Streptococcus bacteria. Then in 2001, as the extent of the threat posed by antibiotic resistance began to emerge, Fischetti gave lysins to mice infected with strep. When the mice were tested shortly thereafter, “Lo and behold,” he says, “the Streptococci were gone!” His experiment became the first in vivo application of phage lysins.
Almost immediately, Fischetti received grants from the Department of Defense to develop lysins against anthrax—a significant national security concern at the time—and other major resistant pathogens. “In those days, every paper we submitted on the topic was quickly published because therapeutic lysins were a new anti-infective,” Fischetti says.
Many laboratories subsequently began pursuing lysins as novel therapeutics, but Fischetti was already a step ahead. One lysin that can fight off a particularly dangerous drug-resistant strain of Staphylococcus aureus was licensed and quickly moved to human trials. It’s the first and only antibiotic alternative to have successfully completed Phase II clinical trials. Fischetti thinks that we could see lysins in the clinic within the next five years.
If so, lysin therapy may prove to be one of the most durable approaches to killing bacteria. “We can’t say it will never occur, but we have yet to see resistance to lysins,” Fischetti says. That’s because the way phages use the enzymes self-selects for success. Phages that produce ineffective lysins never get the chance to pass that defect down to their future generations. Because lysin resistance is so difficult to achieve, lysins could be used to prevent secondary infections. That’s something that antibiotics—as the cause of secondary infections—can’t.
The economic toll of a post-antibiotic age
AGRICULTURE
![]()
63,200 TONS
In 2010, the number of antibiotics that were used in livestock production worldwide, far surpassing human consumption.
In any face-off, the ability to predict your opponent’s next move confers an invaluable advantage. “What if we could predict resistance in the clinic before we brought an antibiotic to market and solve that problem before it gets to the patient?” Brady asks. Each year, the WHO leads a massive international effort to forecast flu strains months out so that effective vaccines can be engineered ahead of the winter season. Is there likewise a way to head off antibiotic resistance at the pass by both better managing how we use existing antibiotics and better understanding how bacteria foil them?
Brady’s most recent work focuses on teasing out the underlying causes of resistance and predicting how quickly and under what circumstances bacteria will become resistant. One day, that kind of information could translate into a ranking system that helps clinicians keep resistance at bay by allowing them to choose the right antibiotic for each circumstance.
But long-term planning and international coordination are key to any successful strategy for combating resistance. As Brady notes, focusing solely on today’s strains in one part of the world doesn’t adequately address the problem; without heading off developing resistance around the globe, scientists can only kick the can down the road.
Consider TB, which generally doesn’t get the same attention (or funding) as resistant staph, strep, and other widespread drug-resistant bacteria “because it doesn’t kill Americans and Europeans nearly as often as it kills people in sub-Saharan Africa and Southeast Asia,” Campbell says. But microbes know no borders, as the pandemic so dramatically illustrated, so addressing multidrug resistance in only one part of the world will ultimately benefit no one. And it turns out that the knowledge generated by studying TB is broadly transferable to other infectious diseases.
Much of Campbell’s work, for instance, centers on unpacking the activities of the TB bacterium’s RNA polymerase (RNAP), the enzyme that creates RNA from a DNA template. But go-to antibiotics invariably target bacterial RNAP in general, meaning Campbell’s efforts shed a broader light on how a range of resistant bacteria dodge frontline antibiotics.
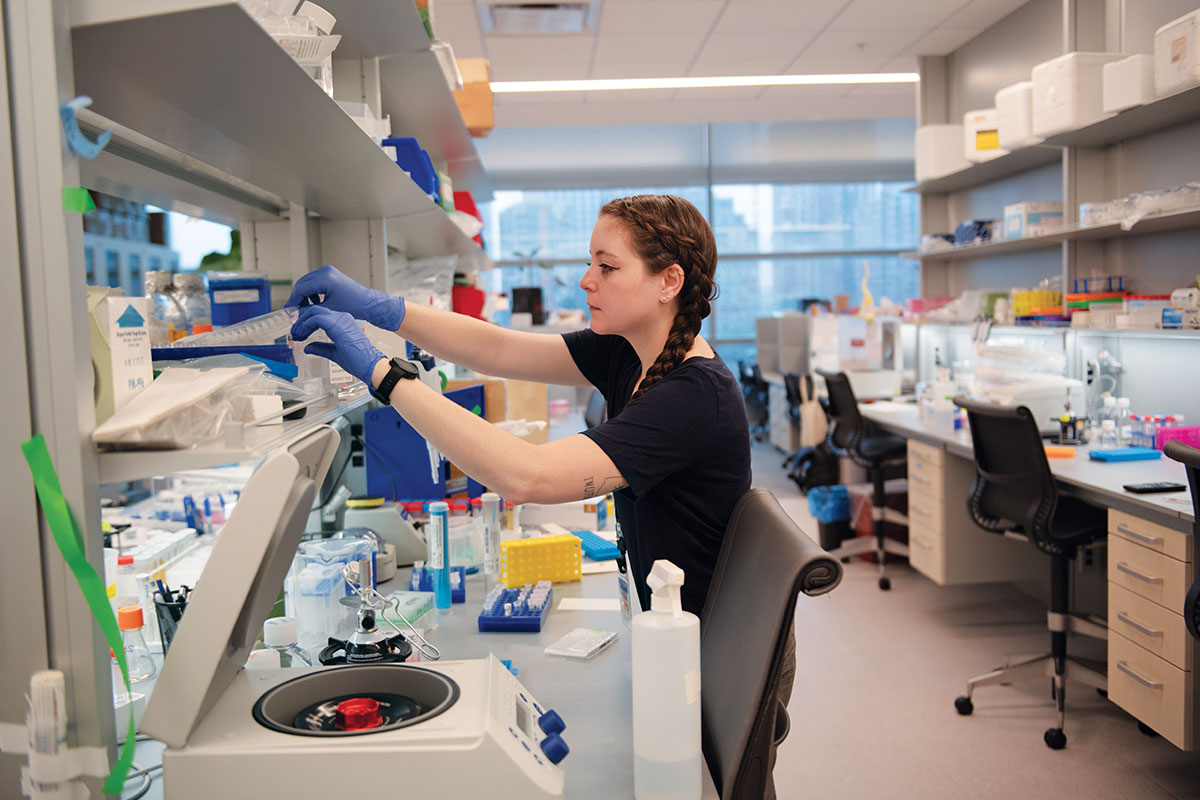

There’s a lot of unsolved fundamental biology here,” says Jeremy M. Rock, the Penrhyn E. Cook Assistant Professor, about the mechanisms that breed antibiotic resistance.
Rock, who heads the Laboratory of Host-Pathogen Biology, is trying to solve those biological mysteries with a large-scale method that lets him knock down almost every gene in Mycobacterium tuberculosis and observe whether that makes the bacterium more or less vulnerable to any given compound. Rock’s lab has used this platform, which employs the revolutionary gene-editing technology known as CRISPR, to discover genes that make TB more sensitive to certain drugs. “You have to wonder,” Rock says, “whether we could use this to rationally design combination therapies for drug-resistant bacteria that would be more potent than expected.”
COVID was a stark reminder of the need to stay ahead of infectious diseases.
What Rock and his colleagues are learning from TB proves the point. Rock and his team have identified 1,373 genes that, when silenced, render the bacteria vulnerable to antibiotics, and another 775 genes that make them more resistant. They found that silencing two genes, mtrA and mtrB, left even resistant bacteria vulnerable to existing therapies and that mutations in a third gene, bacA, may be key to promoting multidrug resistance. They also discovered genes that could allow clinicians to safely use linezolid, an antibiotic that works against drug-resistant TB but only at dangerously high doses. When these genes are inhibited, Rock found that linezolid is effective at far lower—and safer—concentration.
In one of their most promising discoveries thus far, Rock’s team found that a strain responsible for half a million tuberculosis cases each year in Southeast Asia appears to have long ago acquired a mutation that renders the bacteria highly vulnerable to the well-tolerated, FDA-approved family of antibiotics known as macrolides. These drugs have not traditionally been used to treat TB, and deploying them against the disease could be a game changer, especially in the developing world. “Macrolides are cheap, generic, and safe,” Rock says.
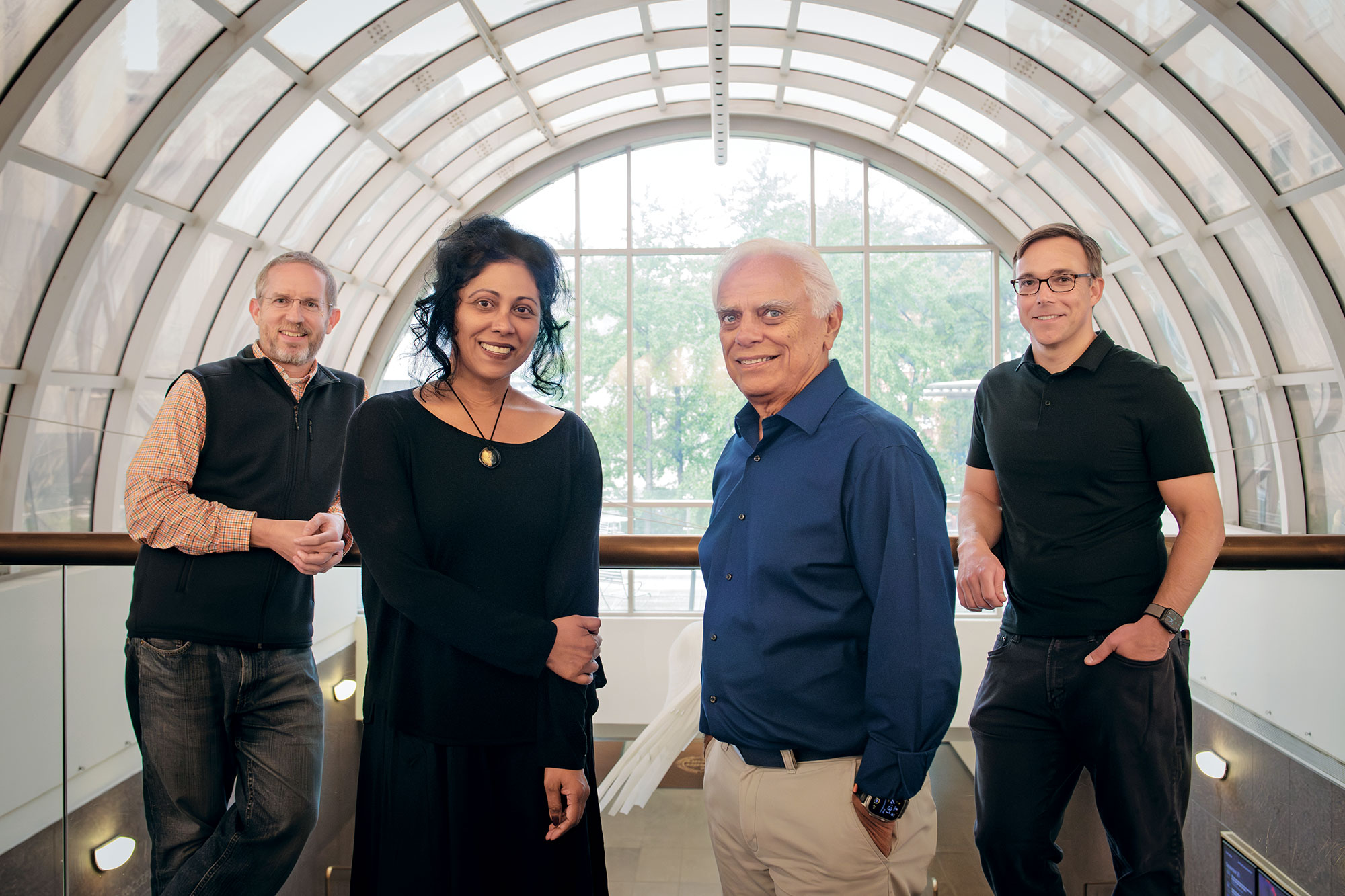

COVID cast a spotlight on how vulnerable we still are to infectious diseases and reminded both those who practice science and those who fund it of the importance of keeping ahead of them. It also illustrated how scientific mountains can be moved when the right resources are focused in the right way.
“What we’re learning in the lab has so much lifesaving potential,” Brady says of the work that he and his colleagues are doing to understand, overcome, and circumvent antibiotic resistance. “But this knowledge has to translate beyond the lab so that we can have alternatives ready to go before we need them.”
With each potential solution—creating novel antibiotics, improving existing ones, or developing entirely new classes of antimicrobial weapons—these scientists’ work is informed by a shared sense of urgency.
Or as Brady puts it, “You don’t build a fire truck after the fire starts.”



