
Interview
The machine that built all living things—and itself
With Sebastian Klinge
Behold the ribosome, enabler of all life, fossil of our primordial past. Colossal among molecules, it has been observed from every angle using a host of elaborate techniques—suffused with electrons, bombarded with X-rays, deep-frozen in liquid ethane. But after decades of dogged research, its myriad nooks and knobs remain blurry.
Until scientists arrive at a perfect picture of the ribosome, their understanding of living things will be incomplete—and so will their ability to heal diseased cells or thwart pathogens.
Discovered at Rockefeller almost 70 years ago, ribosomes have long been known to execute a cell’s most fundamental function: translating its genetic code into protein. But the precise mechanics of that process have been harder to pinpoint. Today, biologists keep adding to the list of “solved structures,” molecules whose 3D shapes have been fully mapped, but the ribosome still holds many secrets. A majestic mountain, it seems too tall to climb.
Sebastian Klinge is not intimidated. He has dedicated his career to understanding how ribosomes string together amino acids, the building blocks of protein, using RNA as a template. Like a watchmaker disassembling an antique timepiece to figure out what makes it tick, he seeks to learn how the ribosome functions by figuring out how one forms. Using ultramodern technology, his lab recently revealed the earliest steps by which numerous ribosome components come together, creating never-before-seen footage of a nonbacterial ribosome folding into shape.
We spoke with Klinge, who heads the Laboratory of Protein and Nucleic Acid Chemistry, about his investigative trek and where it’s headed.
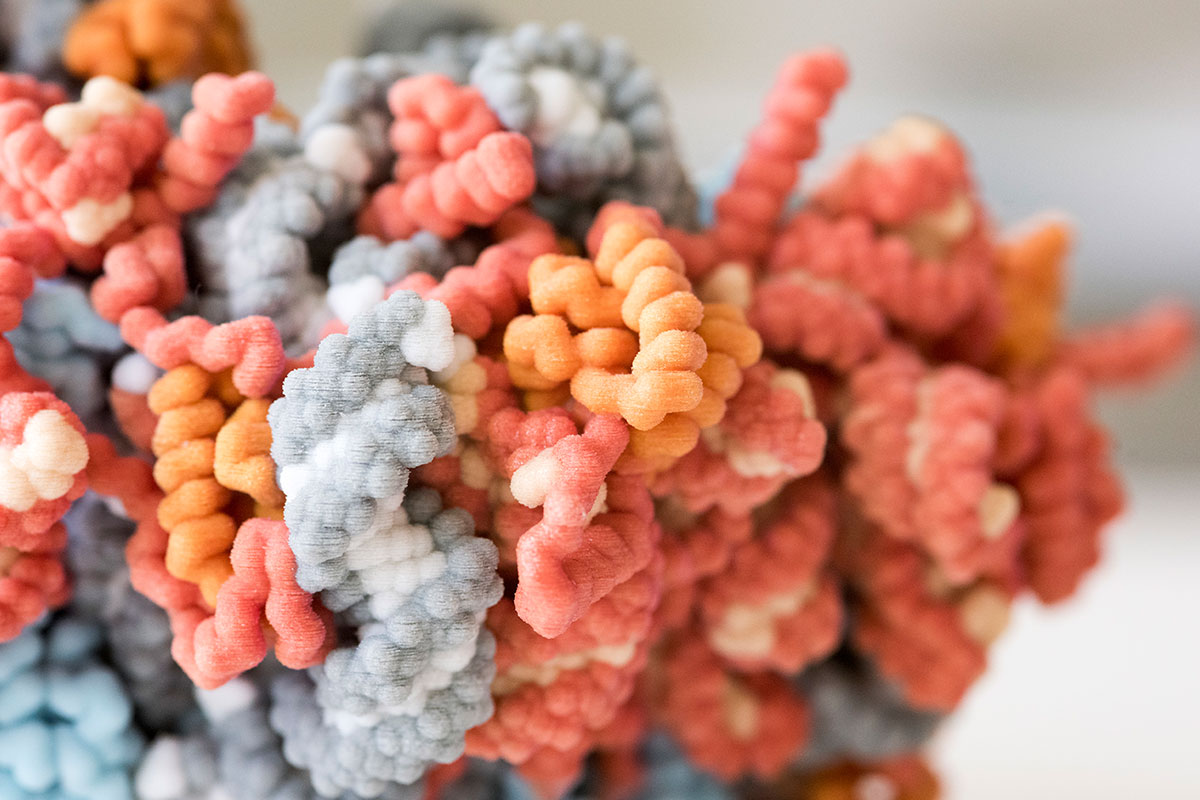
Mapping molecular structures is hard work. Why focus on one as complex as the ribosome?
Ribosomes make the world exist! From an evolutionary standpoint, they have become so indispensable in nature that all cells need ribosomes for everything they do—including making more ribosomes.
Ribosomes are, in fact, so crucial for life that they don’t neatly fit into the boxes in which we generally sort molecules. In most areas of biomedical research, we can tell that a molecule is important by the fact that diseases occur when a mutation arises that prevents it from doing its job. This logic doesn’t apply to ribosomes. If your protein factories aren’t functioning, you’d be unable to even exist. For this reason, we rarely see cells or organisms with serious mutations in the ribosome. And the minor ribosomal mutations that don’t immediately kill the organism tend to have major consequences, causing cancer predisposition or craniofacial malformations.
If cells need ribosomes to make ribosomes, where did the first ribosome come from?
It’s almost a chicken-and-egg problem, because that’s right—to make a ribosome you need RNA as well as ribosomal proteins, which would have been made by another ribosome.
There must have been a point in evolutionary history when the ribosome was much simpler, a machine consisting only of RNA that manufactured polypeptides. In all likelihood, some of those polypeptides ended up interacting with the RNA and so thoroughly stabilized it that they eventually became part of the machine itself. Over time, these proteins evolved to become more efficient and sophisticated—and better at avoiding errors—eventually becoming ribosomes as we know them today.
How much do we know about how ribosomes are put together?
For a long time, we didn’t even know what ribosomes looked like. When Rockefeller scientist George E. Palade first discovered them in 1955, he called them “particulate components of the cytoplasm” because they appeared as dark dots under the electron microscope. Decades later, in the late 1990s, X-ray crystallography started to reveal the many different components of ribosomes, yet we still didn’t know how these structures were assembled.
“It’s a chicken-and-egg problem:
You need ribsomes to make ribosomes”
Imagine archaeologists unearthing an ancient structure but having no idea what materials it was made of or how it had been put together. This was our problem. We had detailed 3D images of the ribosome, but still no idea how it was built. Only in recent years has the development of cryo-electron microscopy technology enabled my lab and others to gain insight into the ribosome assembly process.
Even cryo-EM only yields a snapshot, though. We’re still trying to string snapshots of ribosomal formation and activity together into the right order to understand how the ribosome gets from one point in its development to the next—no easy task with molecules at this level of complexity. We’ve only recently accumulated enough molecular snapshots of these pathways that we can begin to tackle this task, putting together movies that allow us to really understand how ribosomes form and operate.
How might this work inform our understanding of human disease?
Although the vast majority of ribosomal mutations are so deleterious that they cannot sustain life, we know of some that are subtle enough to cause disease without killing the organism. Specifically, a handful of blood diseases have been linked to mutations in ribosomal proteins.
Research has shown that mutations in ribosomal proteins can decrease the amount of ribosome assembly taking place or the quantity of ribosomal proteins available to synthesize a transcription factor critical to the development of red blood cells. The upshot is that the body is unable to produce sufficient red blood cells and their overall count drops, leading to anemia and related problems. Once we have a better grasp of how ribosome assembly happens, and how it fails, we might have opportunities to develop new therapies to manage these conditions.
Data
Each of the body’s cells may contain up to 10 million ribosomes.
A better understanding of ribosome assembly may also lead to novel therapies for microbial disease. Many existing drugs kill bacteria or fungi by targeting their ribosomes. By studying ribosomal structures, we have been able to pinpoint the precise mechanisms by which some of these drugs work. For example, we now know where the antifungal cycloheximide binds in eukaryotic cells and where the antibiotic azithromycin binds in bacterial cells.
Ultimately, this research might aid the discovery of new antimicrobial treatments with better specificity and fewer side effects—drugs capable of attacking the pathogen without destroying beneficial bacteria or human cells.
What else about ribosomes do we still need to study?
Now that we are beginning to understand the principles of the ribosome, the basic machinery, and its parts, we can move on to study how ribosomes are controlled and regulated. For instance, how does a cell decide which messenger RNAs should get translated into a protein? How does the ribosome make new ribosomes? And how does cellular metabolism influence all of this? One crucial question that we’re looking into right now is how the control and integration of ribosomes works. We still don’t know how misassembled ribosomes are recognized and what controls ensure that ribosomes are assembled correctly.
These questions are the future of ribosome research. At Rockefeller, where we have a rich history of ribosome discovery and investigation, I am just the latest in a long line of people working on these problems.